PoET-2’s architecture generalizes to families and properties it has never seen.
Protein language models capture evolutionary patterns shaping protein structure and function
Developed by the pioneers of protein language models
Sign UpGenerate diverse, functional sequences de novo given sequence and structural constraints
Learn more →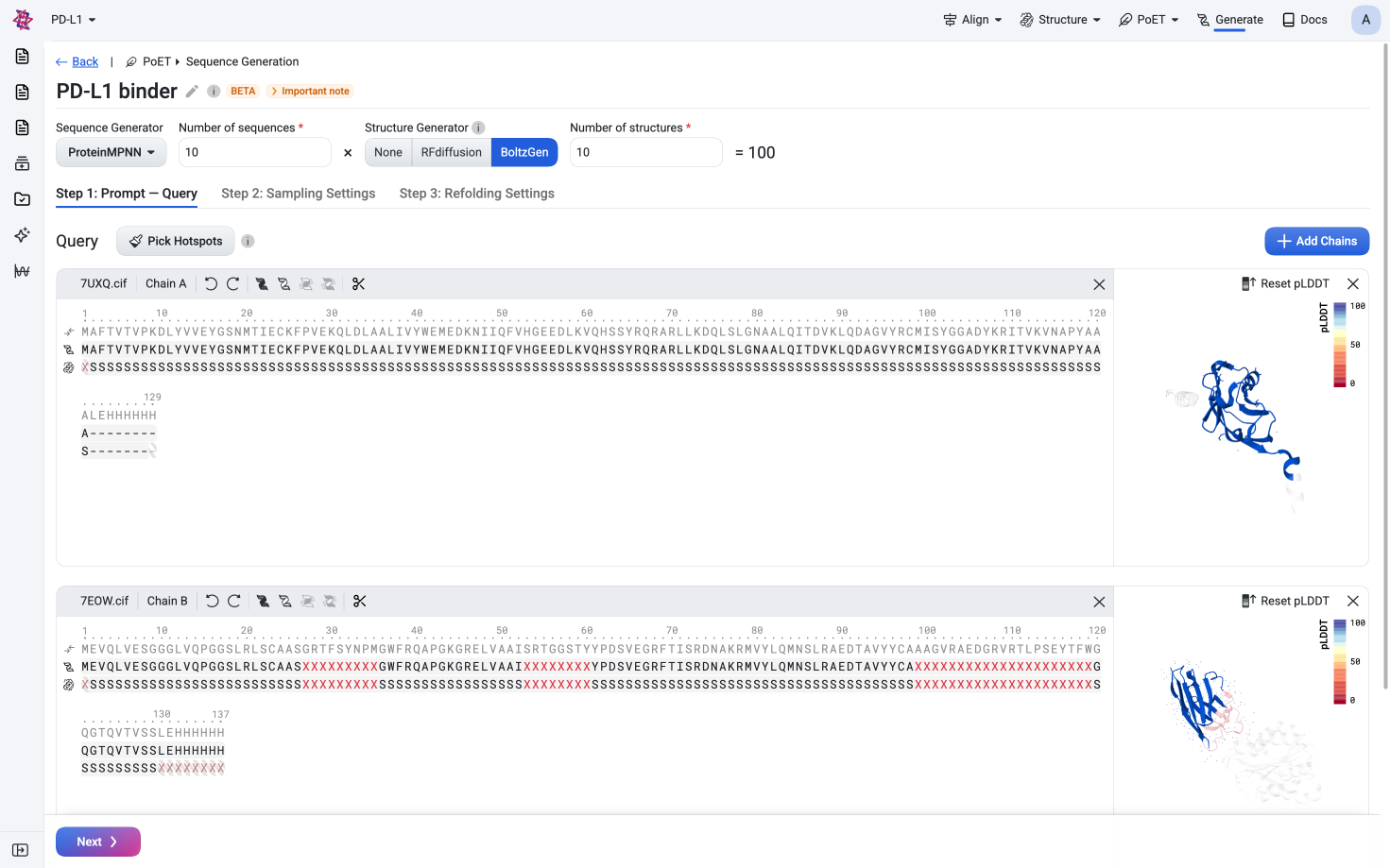
Learning from billions of years of evolution
...to drive end-to-end protein engineering
Protein language models capture evolutionary patterns shaping protein structure and function
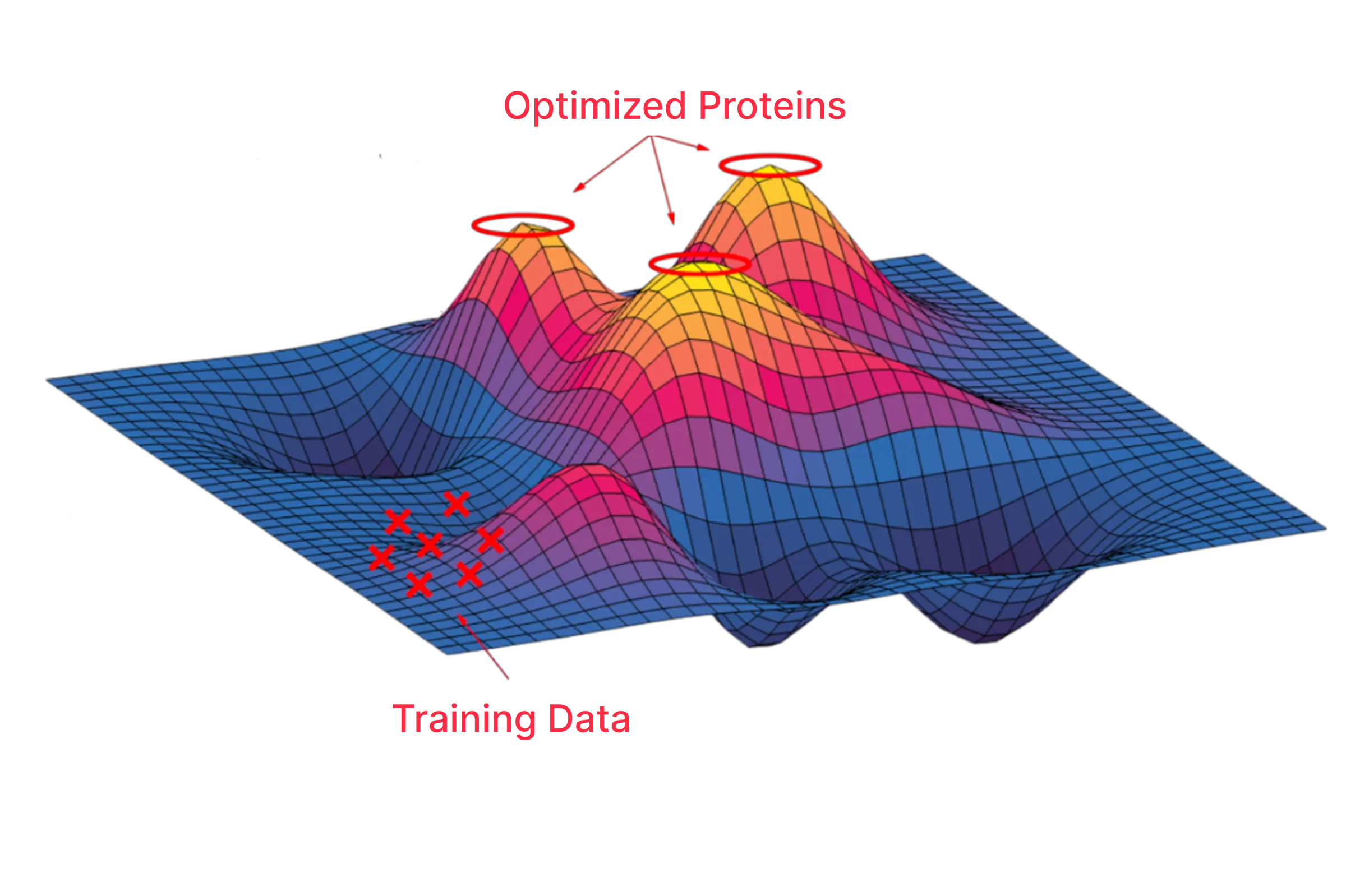
Design protein sequences de novo
Analyze the fitness landscape and prioritize variants
Zero shot variant effect prediction
Evaluate the functional impact of novel variants in silico - including indels
Learn from your mutagenesis data
Train specialized models on your own data
Screen libraries with specialized models
Rank and filter sequences to focus on the most promising candidates
Design combinatorial variant libraries
Optimize variants for multiple properties
Generate sequences
Variant effect prediction
Train models
Predict variants
Design libraries
From de novo design to optimization, OpenProtein.AI provides the tools and AI infrastructure for protein design. Accelerate your design-build-test cycles and maximize the value of every data point.

Reduce costs more than 60%

Accelerate your projects more than 3x

Eliminate guesswork
End-to-end protein engineering suite, easily accessible through a powerful graphic interface.
Programmatic access to models for design, optimization, and analysis. Built to scale.
Integrate models, data systems, and ML frameworks across experimental and computational teams
Directly leverage ML advances
Accelerate research with self-service, no-code access to your team’s ML models, instantly integrated directly into existing workflows
Deploy custom models directly to biology teams
Build better models faster with managed infrastructure, and seamless deployment to biology teams
State-of-the-art model for protein design
Pushing the boundaries of protein language models
PoET-2’s architecture generalizes to families and properties it has never seen.
State-of-the-art zero-shot performance across benchmarks spanning fitness, stability and binding. Design protein variants with no assay data required.
Train more accurate sequence-property predictors with 10x less data, cutting assay costs and accelerating iteration cycles.
Access protein foundation and generative models all in one place. Use these models to design new proteins, fine-tune them with new data, or run in silico screening with our APIs or GUI. Compare between models to find the best model for your application.
Embed, score, and generate sequences conditioned on a protein family. PoET-2 also accepts sequence and structural inputs as constraints and context.
Encoder-only transformer models trained on large protein sequence datasets. Supports embedding, scoring, and attention maps.
Citation: Lin, et al., Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123-1130 (2023). doi:10.1126/science.ade2574
Encoder-decoder transformer models trained on large protein sequence datasets. Primarily used for embedding and representation learning.
Diffusion model for novel backbone and complex structure design. Supports motif scaffolding and binder design.
Citation: Watson, et al., De novo design of protein structure and function with RFdiffusion. Nature 620, 1089–1100 (2023). doi:10.1038/s41586-023-06415-8
Generates 3D structures and conformational ensembles conditioned on sequence, motifs, or interaction constraints.
Citation: Stark, et al. Toward universal binder design with BoltzGen. bioRxiv 2025.11.20.689494 (2025). doi:10.1101/2025.11.20.689494
Predict static and dynamic biomolecular structures and binding affinity to support structure-based drug discovery.
Citation: Passaro, et al. Boltz-2: towards accurate and efficient binding affinity prediction. bioRxiv 2025.06.14.659707 (2025). doi:10.1101/2025.06.14.659707
Predicts 3D protein structures from amino acid sequences using deep learning and evolutionary information.
Citation: Jumper, et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021). doi:10.1038/s41586-021-03819-2
Predicts 3D protein structures directly from sequences using transformer-based language models trained on large-scale datasets.
Citation: Lin, et al. Language models of protein sequences at the scale of evolution enable accurate structure prediction. Science (2023). doi:10.1126/science.ade2574
Lightweight transformer for fast embedding, scoring, and representation learning on downstream ML tasks.
Citation: Chen, et al. Protenix: advancing structure prediction through a comprehensive AlphaFold3 reproduction. bioRxiv 2025.01.08.631967 (2025). doi:10.1101/2025.01.08.631967
Compact AlphaFold-style architecture for rapid structure predictions with reduced compute on small–medium proteins.
Citation: Alcaide, E. (2019). MiniFold: a DeepLearning-based Mini Protein Folding Engine. GitHub. doi:10.5281/zenodo.3774491
Integrates sequence, structure, and diffusion-based reasoning for improved complex prediction and conditional design.
Citation: Baek, et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876 (2021). doi:10.1126/science.abj8754
...and more
Understanding protein function with a multimodal retrieval-augmented foundation model
Timothy F. Truong Jr, Tristan Bepler
NeurIPS 2025
Machine learning optimization of candidate antibody yields highly diverse sub-nanomolar affinity antibody libraries
Lin Li, et al.
Nature Communications 2023
PoET: A generative model of protein families as sequences-of-sequences
Timothy F. Truong Jr, Tristan Bepler
NeurIPS 2023
Combine proprietary data with foundation models to train specialized models tailored to your needs. Deploy these models to accelerate new projects.
Sign up now →
No usage barriers - transparent
pricing
Work with the top experts in protein ML

Plug and play system - no setup required
Best practices workflows out of the box
Connects experimental and computational teams
We safeguard your data with encryption, full account isolation, and secure cloud infrastructure. No one accesses your data but you.
Retain full IP rights to your dataset, designs and output. No royalties, no licensing fees.
Accelerate protein engineering and unlock the full potential of your data
Sign Up Now