Data-driven Protein Engineering
Discover functional protein sequences optimized to your specifications
Get Early AccessIntuitive and accessible web app
Machine Learning-driven protein engineering at your fingertips
Deploy state-of-the-art ML models based on your sequence and function data to generate new, more diverse variants. No specialized skill required.
Machine learning-guided mutagenesis
Powerful analytical tools to increase your success rate over standard mutagenesis
The OpenProtein.AI web app provides a suite of software tools to generate novel variant libraries and predict their success over multiple functions of interest. Visualize your mutagenesis data, train machine learning models for functions of interest, define your design objectives, and build optimized variant libraries.
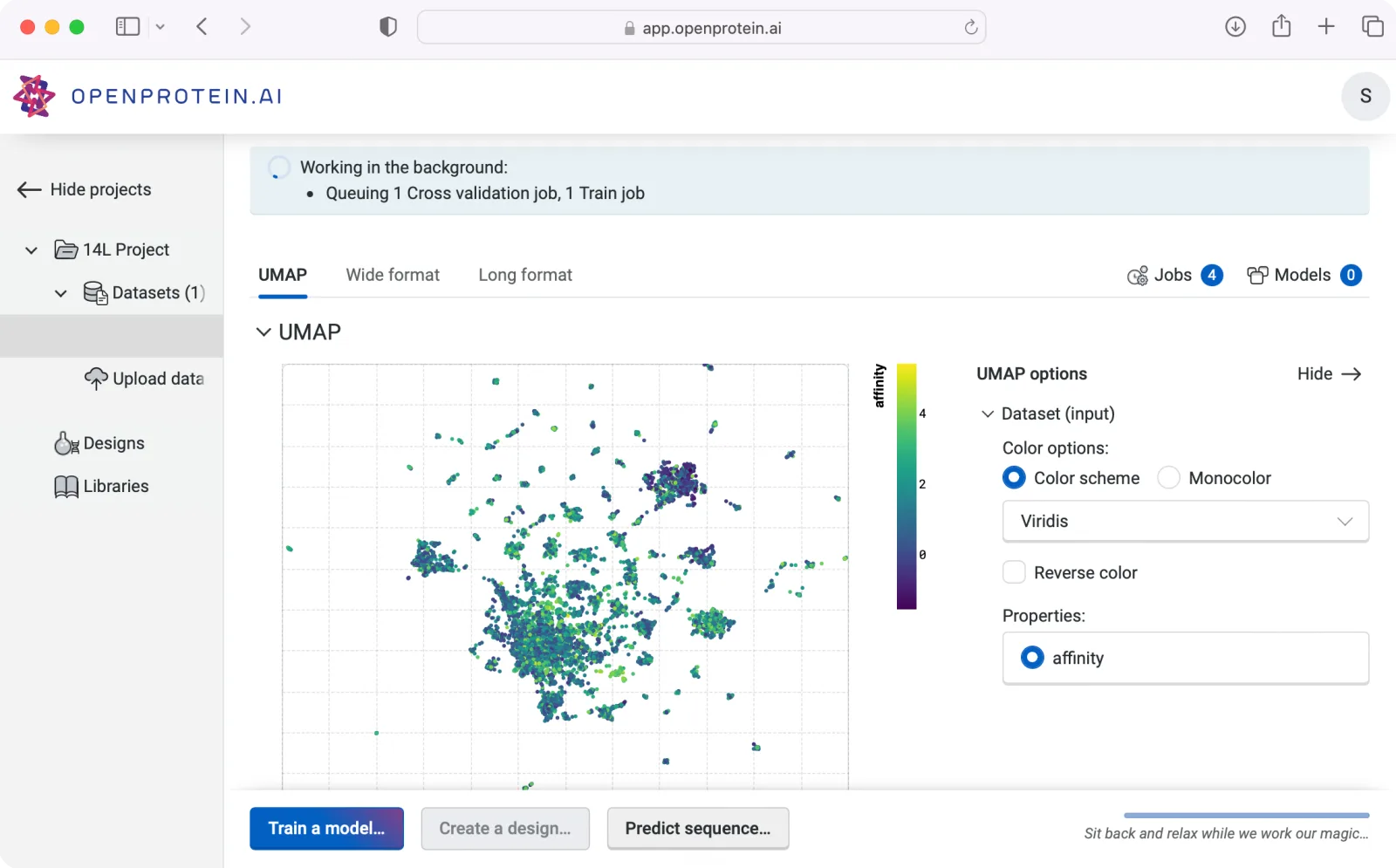
Convenient, reliable data management
Track your mutagenesis process and manage your data all in one place
Streamline your research process with advanced in-app data management capabilities. OpenProtein.AI is a secure data repository for large mutagenesis datasets.
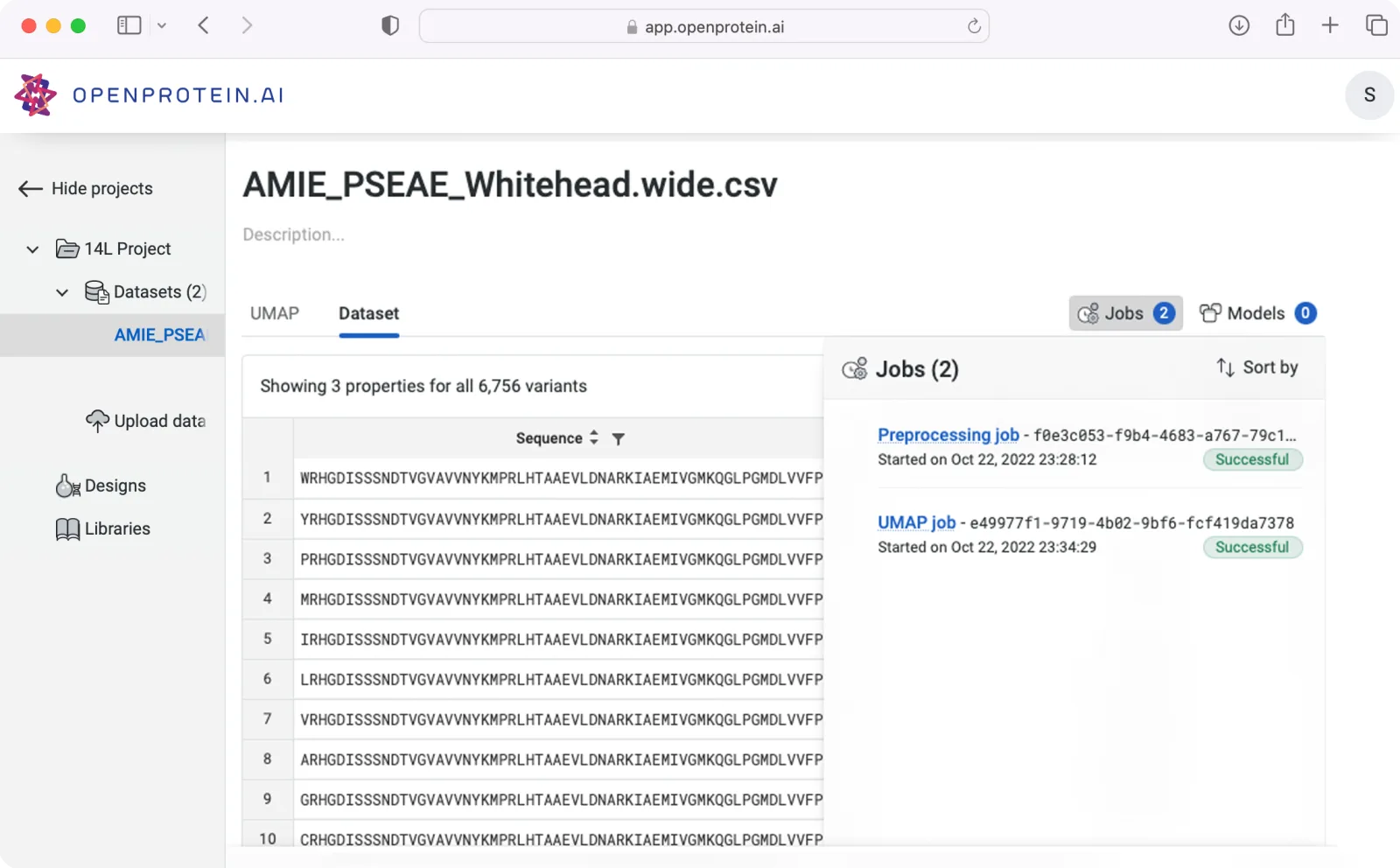
Data-driven protein engineering
Unlock your data's full potential
OpenProtein.AI mines natural sequence databases and learns from your experimental data to accelerate the iterative design process. Design variants with significantly enhanced activity compared to standard directed mutagenesis.
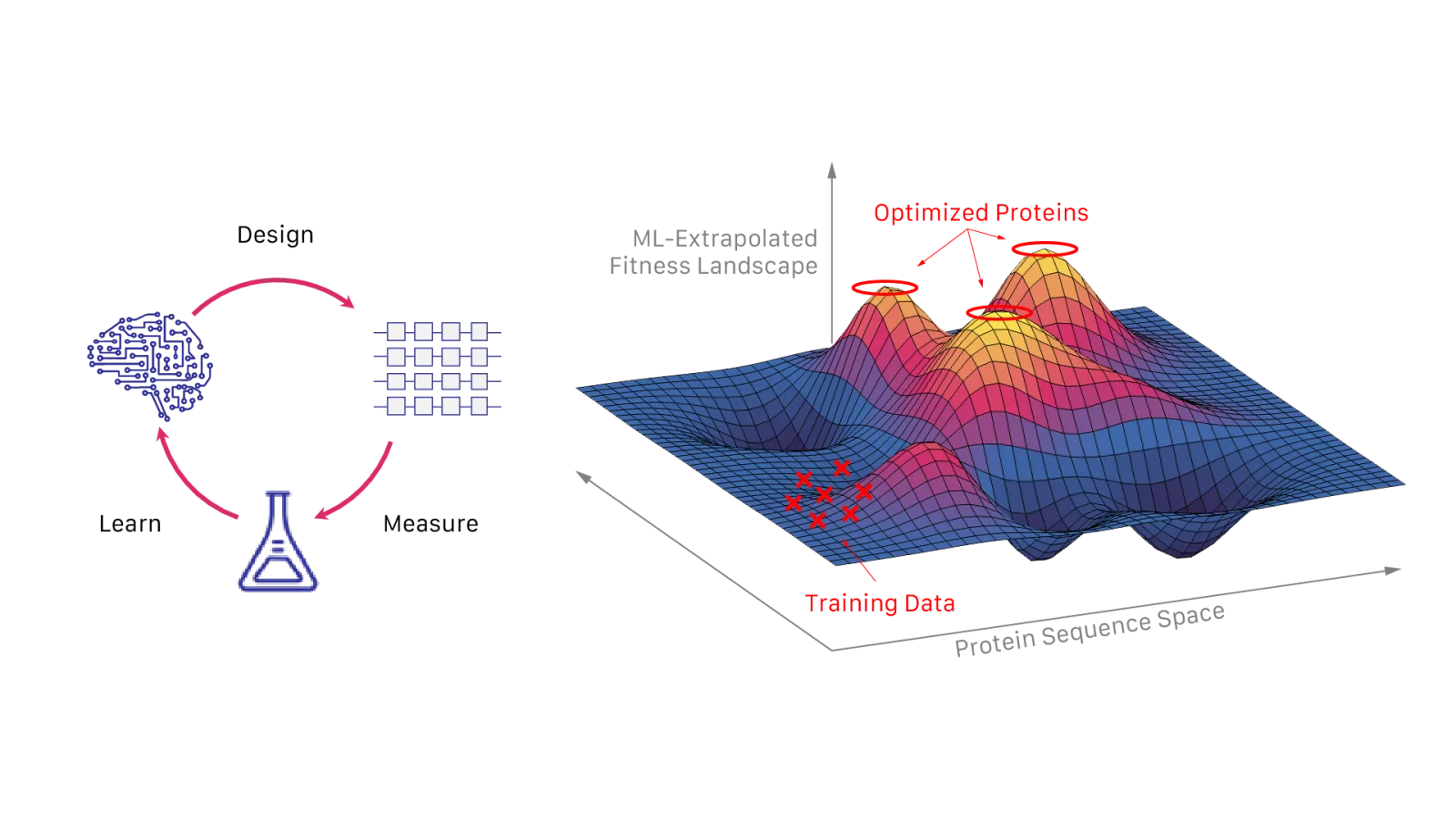
Experimental efficiency
Optimize multiple properties simultaneously
OpenProtein.AI can improve multiple properties simultaneously to reduce experimental iterations. Every subsequent round and project benefits from previous data.
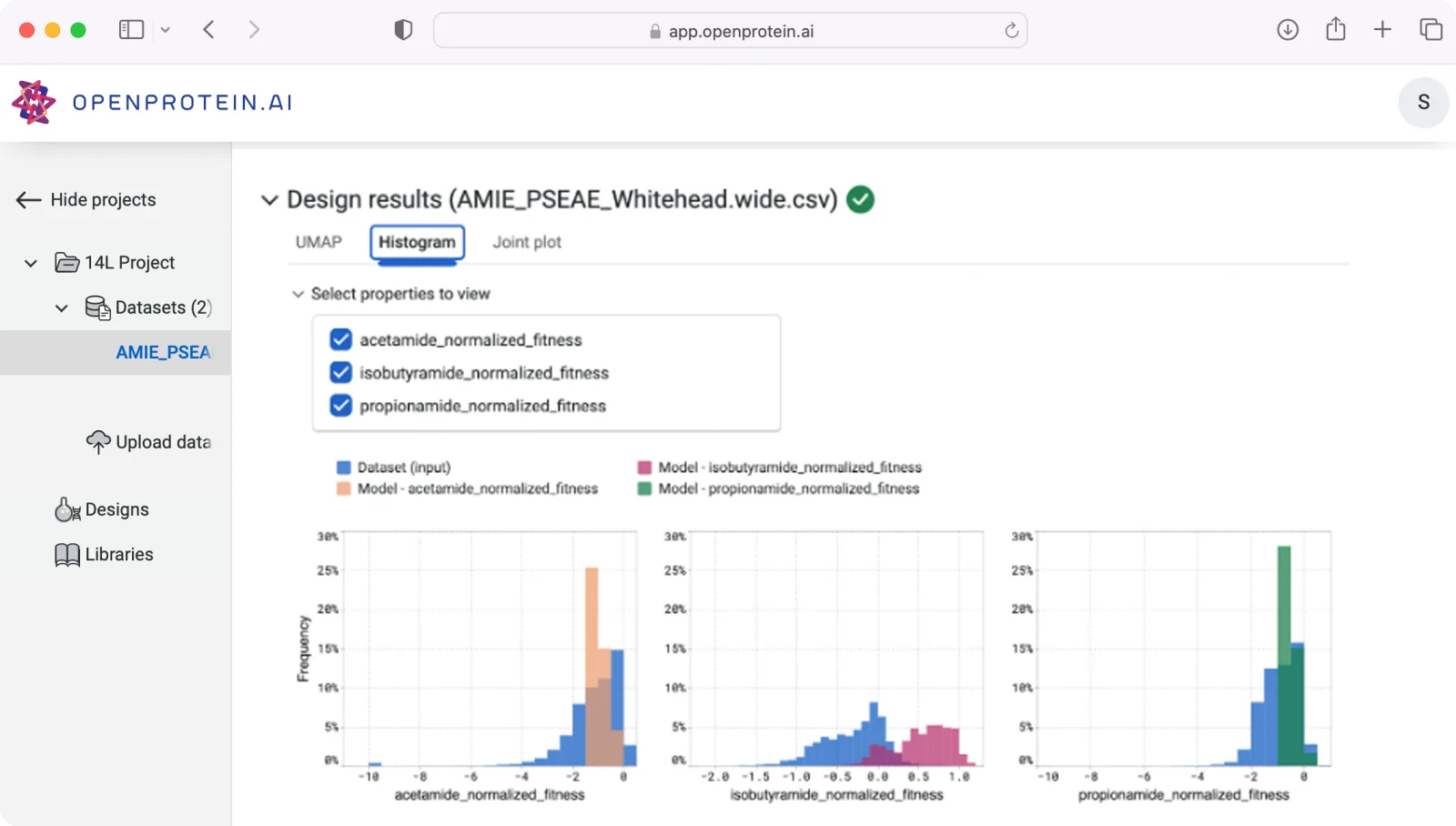
Sequence-to-function mapping
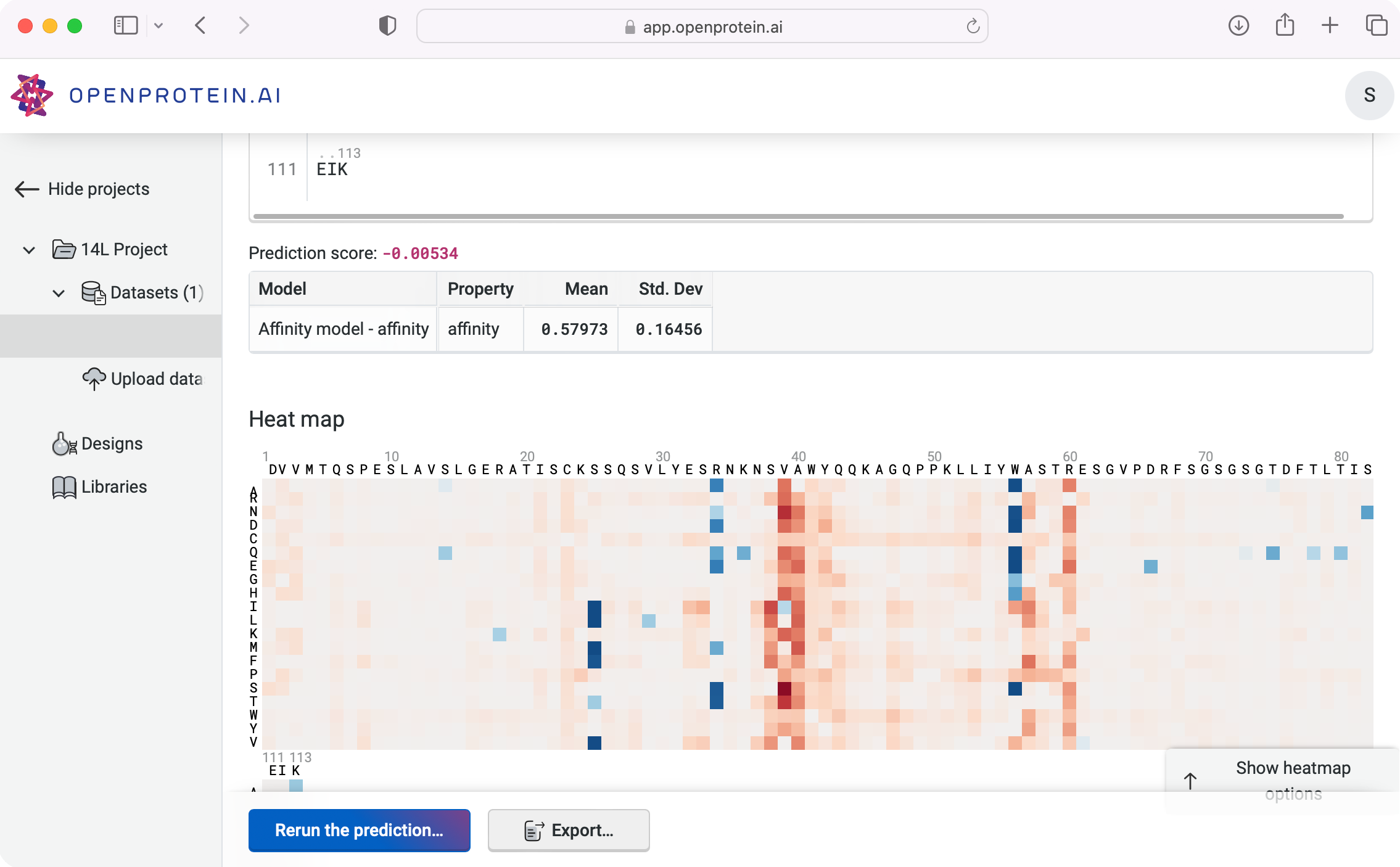
Predict functions of interest, identify mutagenesis hotspots, and design combinatorial variant libraries
Develop & deploy models based on your data to predict activity for any input sequence and map all single site substitutions to identify linchpin locations for site-saturating mutagenesis. Visualize functional predictions for all single-site substitutions and export amino acid distributions for degenerate and combinatorial variant libraries.
Powered by AI. Inspired by evolution
Generative protein design with PoET
Design protein sequences de novo, no functional or structural data required
Sign UpFree for academic use!
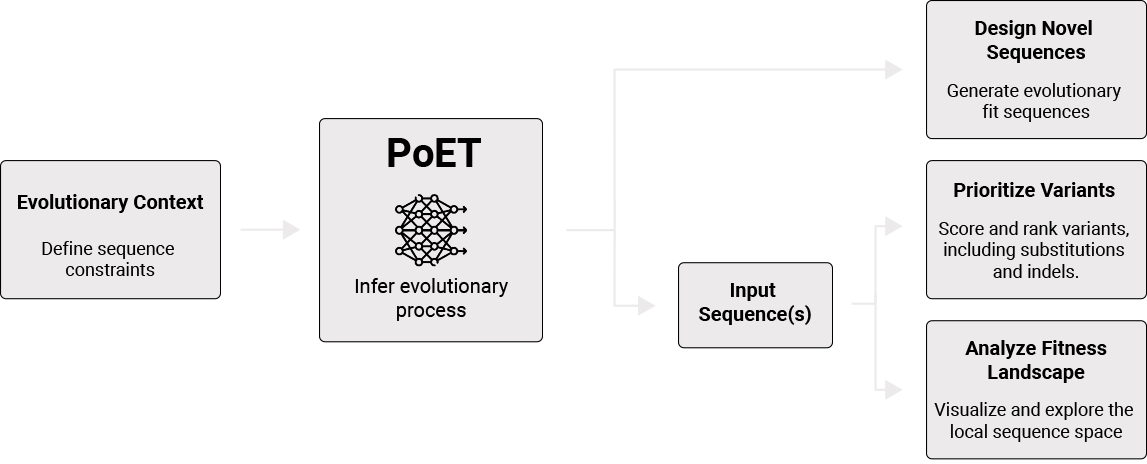
What is PoET?
PoET (Protein Evolutionary Transformer) is an autoregressive retrieval-augmented generative transformer protein language model.
Given a set of sequences representing the evolutionary context, PoET directly infers the fitness landscape on which natural selection acts to optimize proteins under functional constraints on the amino acid sequences. PoET can then generate new sequences from that evolutionary process or score the fitness of arbitrary query sequences under that process.
Generate novel, functional, and diverse sequences
PoET efficient sampling from the learned evolutionary process
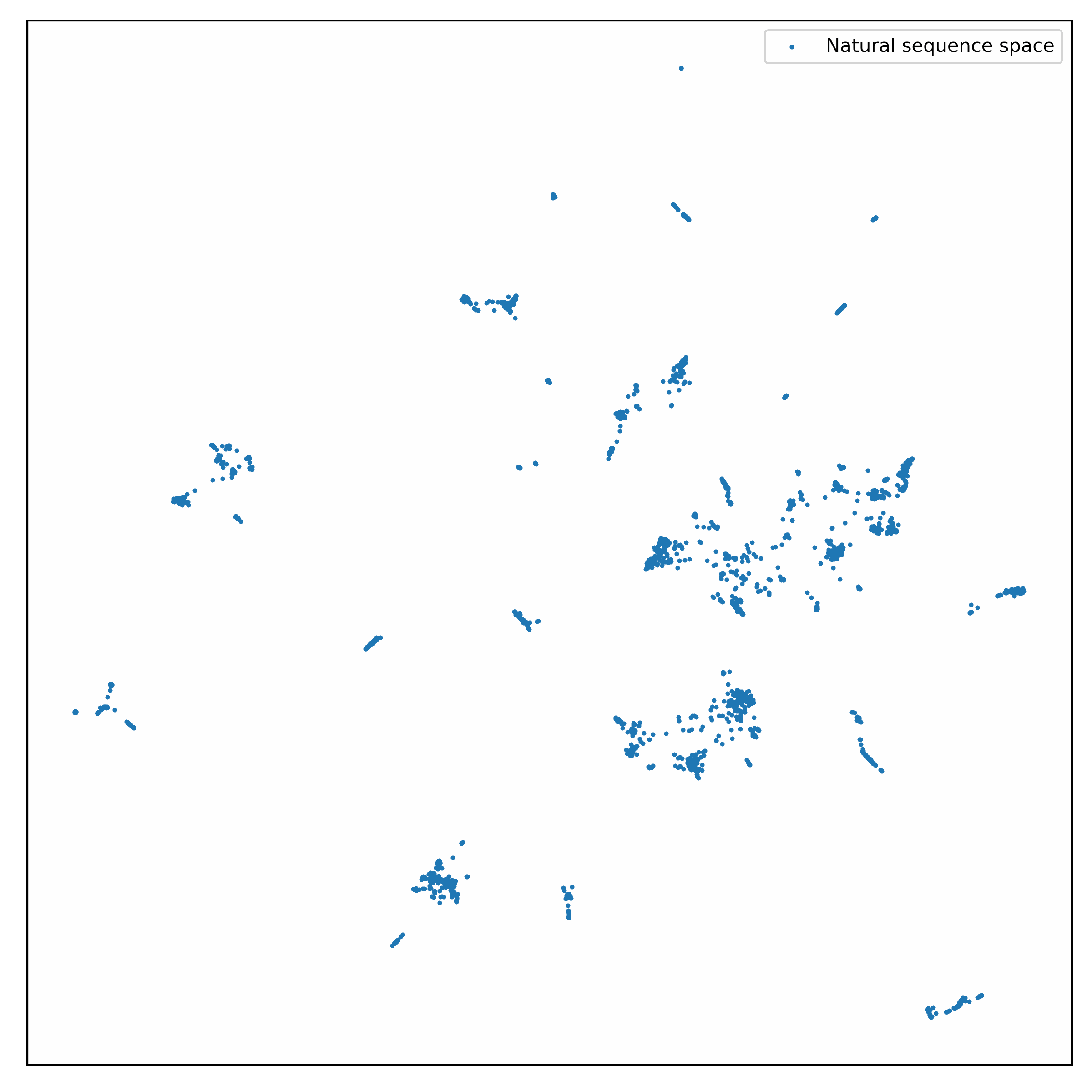
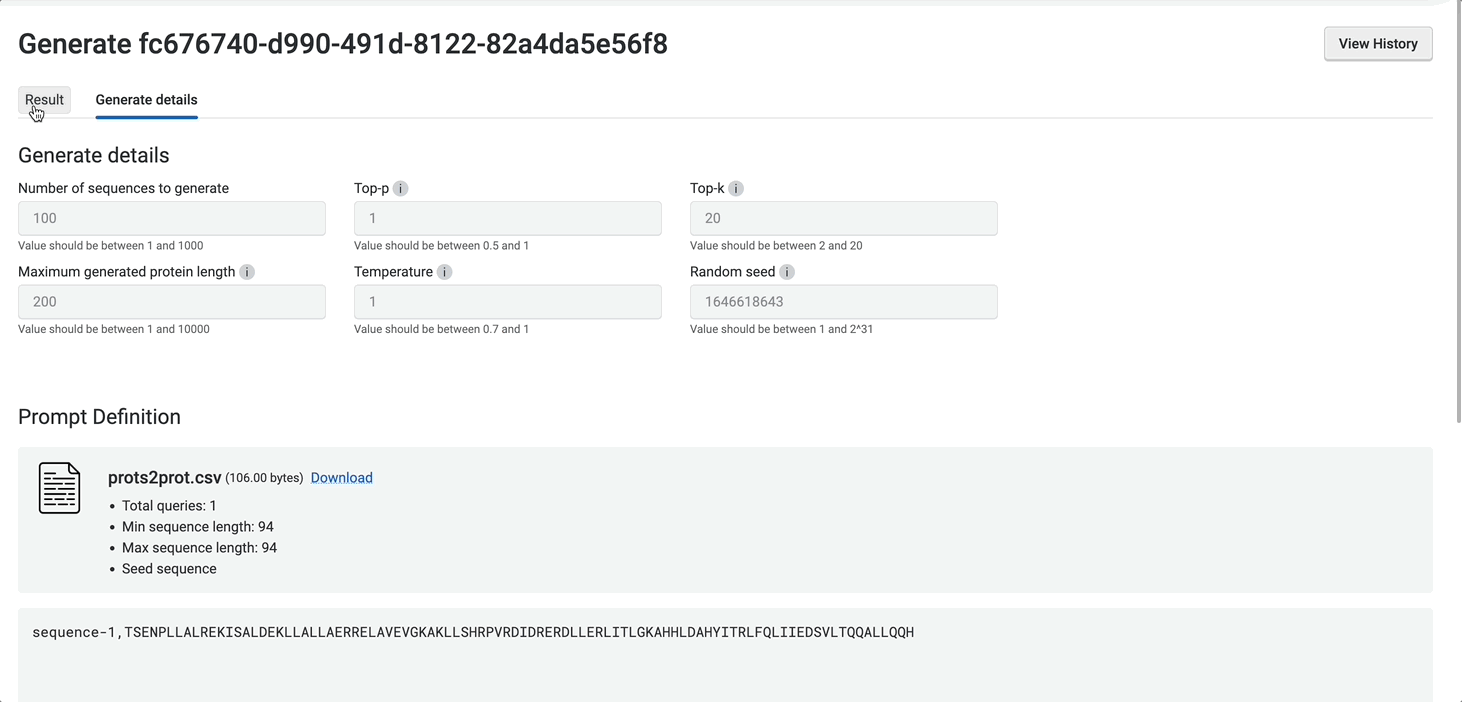

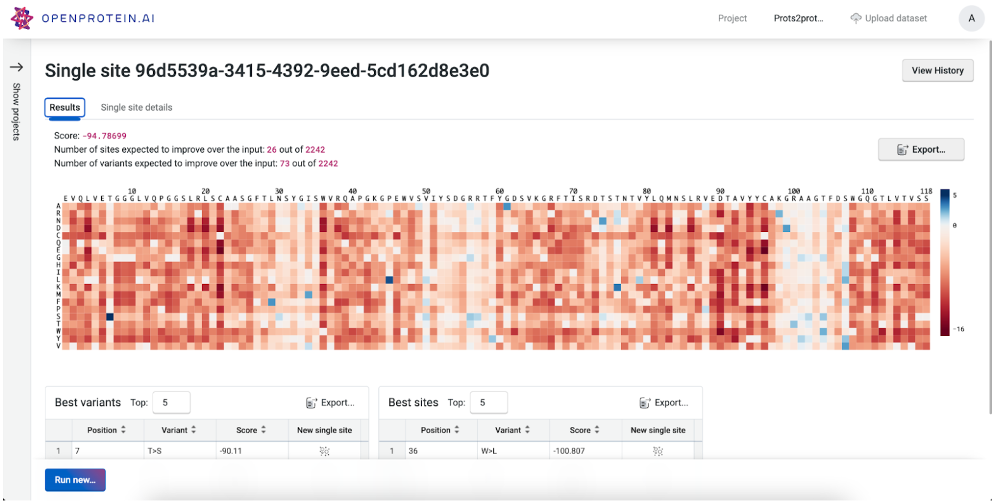
Analyze the fitness landscape and prioritize variants
Given a parent sequence, explore the local fitness landscape or rank specific variants to designs focused mutagenesis libraries
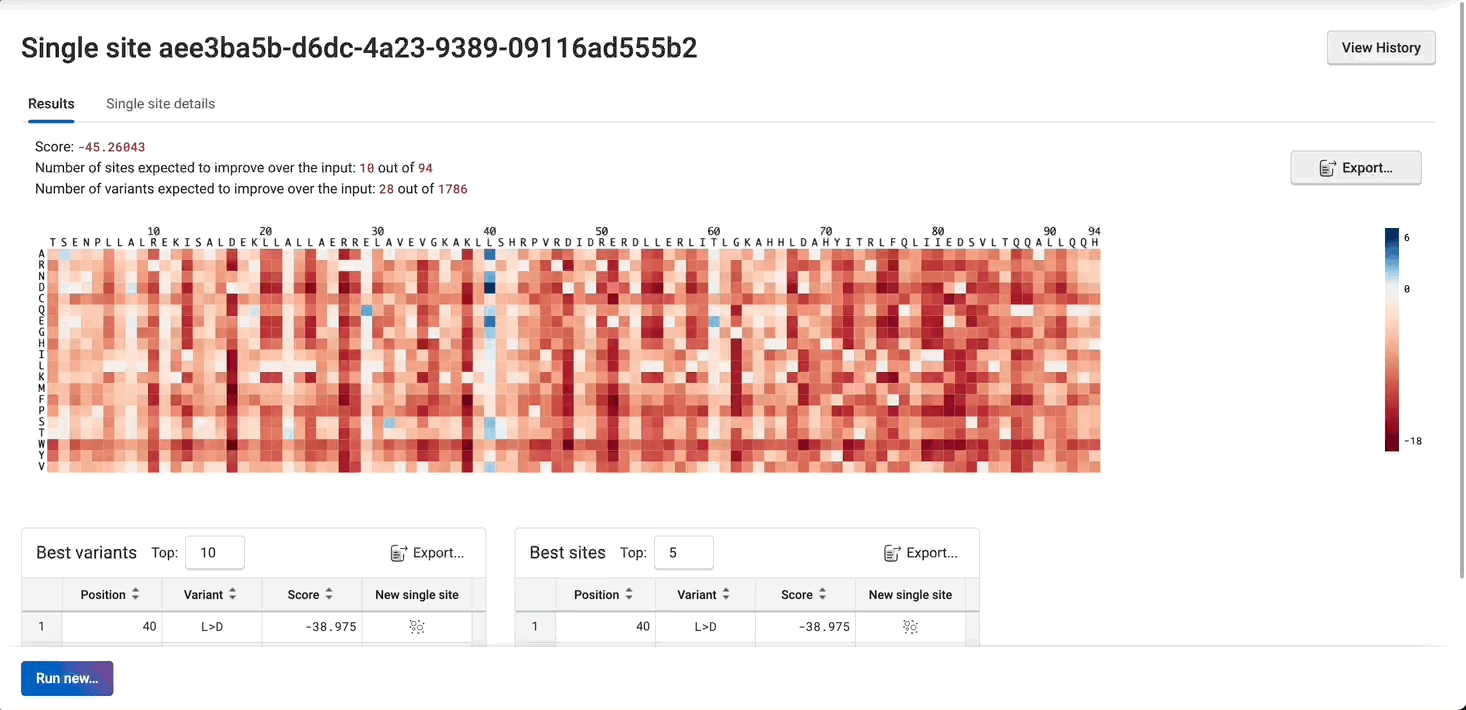
Sequence-to-function mapping
PoET is simple to use and works out of the box
Intuitive workflows are quick and easy to use. Results are returned in minutes and can be exported in multiple formats.
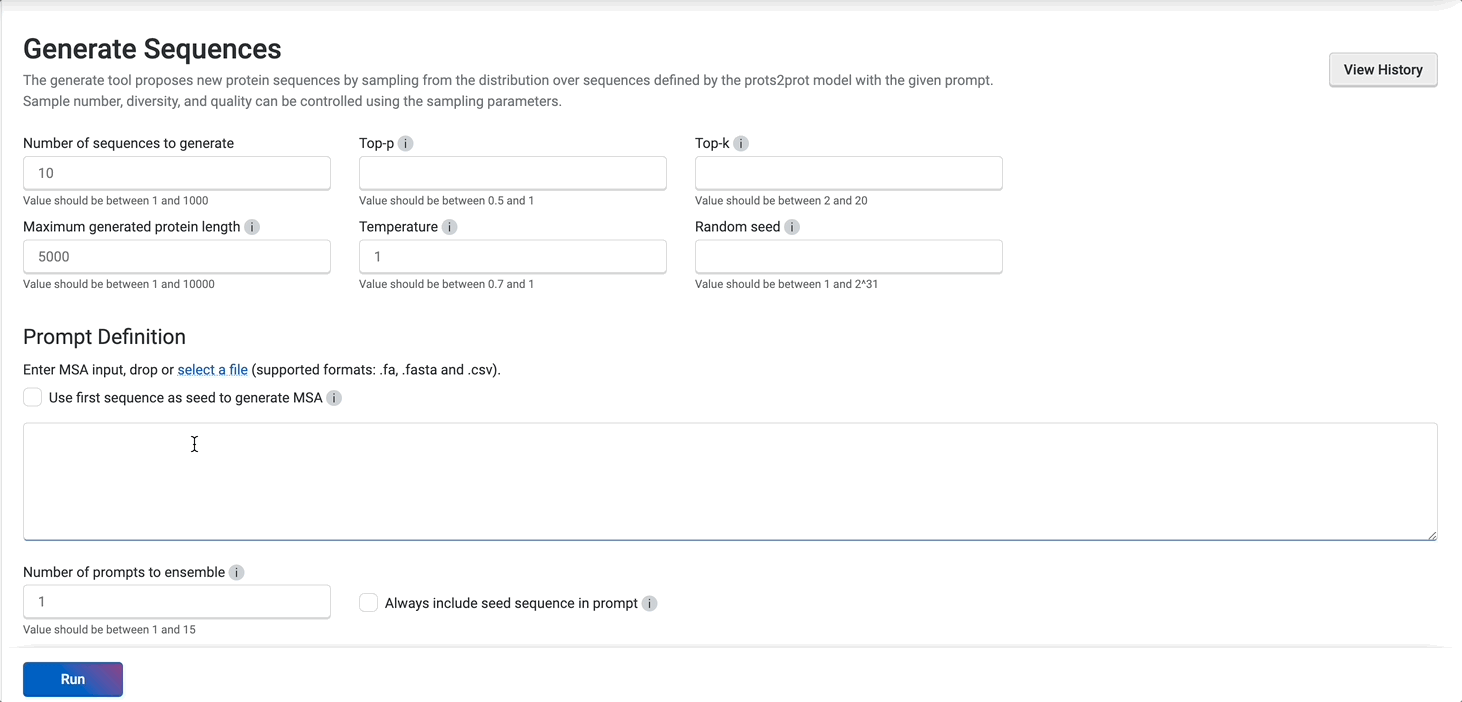
Tailor your designs
Specialize PoET to your applications
Define your evolutionary context through prompt customization. Use any sequence database with custom MSAs. Adjust diversity of the model with in-software similarity level settings.
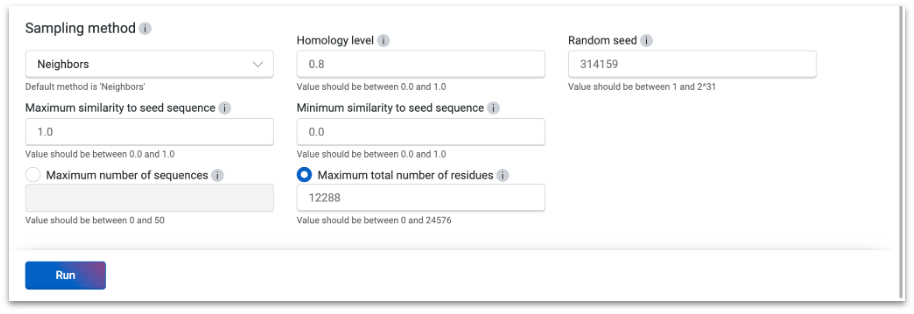
State-of-the-art variant effect prediction
Validated on 90 different deep mutational scanning datasets
PoET provides state-of-the-art de novo variant function predictions across a wide range of
- protein families,
- organisms of origin,
- properties of interest, and
- MSA depths.
PoET can model
- substitutions, insertions, and deletions,
- single and higher order variants.
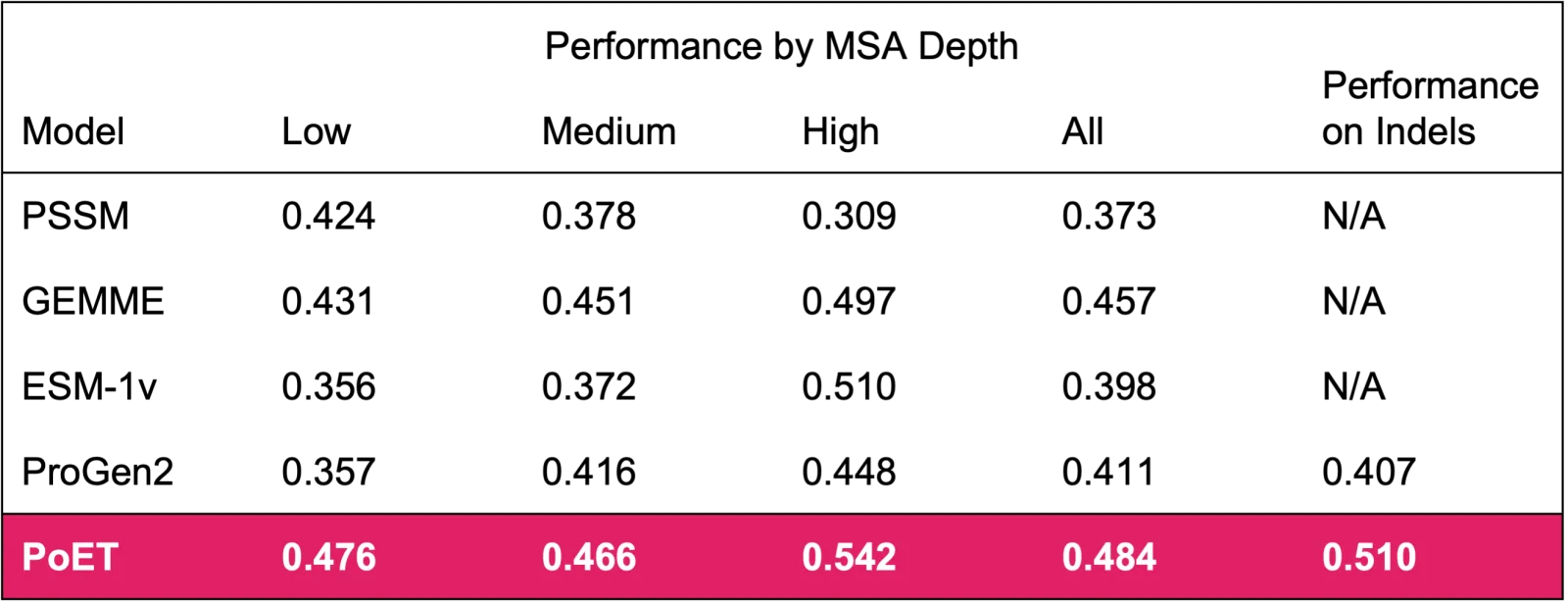
Performance is measured as the rank correlation between variant likelihoods and measured function. N/A is reported for models that cannot predict indels.
Enhanced mutagenesis workflow
Engineer better proteins, faster!
Variant Library Design Features
- Evolutionary sequence analysis
- Generative protein language models
- Identify mutagenesis hot spots
- Design combinatorial variant libraries
- Optimize variant libraries for multiple design objectives
Variant Fitness Predictions
- Train models to predict function(s) from your mutagenesis data
- Predict variant sequence activity for functions of interest
- Perform single site substitution, deletion, and insertion analyses
- Create likelihood-activity relationship generative models
Actionable Results
- Identify target substitution, insertion, and deletion sites
- Design single or higher order variants with enhanced activity
- With statistical coupling analysis, discover areas with high potential for epistasis